Just a quick tip. Sometimes I have a column in a Vortex workspace that I want to access via script where the title is something
Tag: vortex
UniChem is a web resource provided by the EBI, it is a ‘Unified Chemical Identifier’ system, designed to assist in the rapid cross-referencing of chemical structures,
UniChem is a web resource provided by the EBI, it is a ‘Unified Chemical Identifier’ system, designed to assist in the rapid cross-referencing of chemical structures,
Promiscuous inhibition caused by small molecule aggregation is a major source of false positive results in high-throughput screening. To mitigate this, use of a nonionic
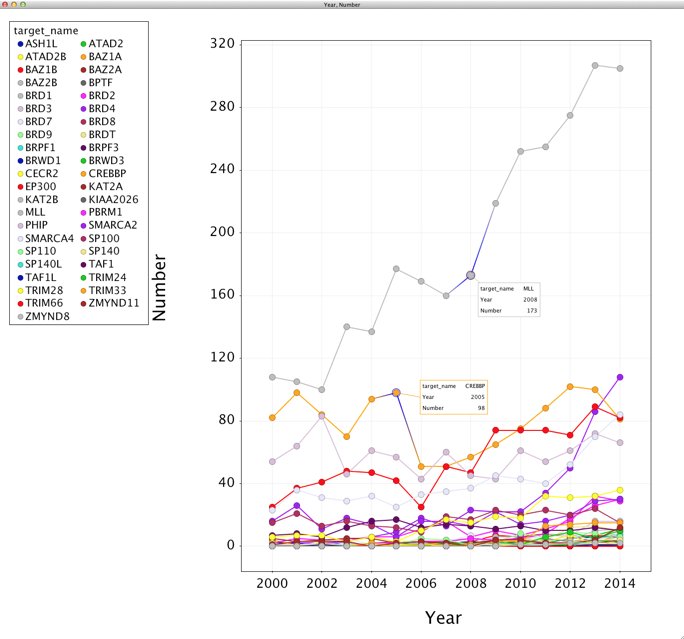
Whilst most of the Vortex scripts mentioned on this site to date involve chemical structures we should not forget that Vortex is an excellent general
In an earlier script I described the use of the ability to script multiple sub-structure searches using SMARTS. There are many occasions when this sort

ChEMBL is a manually curated chemical database of bioactive molecules . It is maintained by the European Bioinformatics Institute (EBI), of the European Molecular Biology Laboratory
One of the really neat features of the latest version of Vortex (> build 29622) is the ability to script multiple sub-structure searches using SMARTS.
Well things can change quickly at times, in the last tutorial I wrote.. Vortex has a limited capacity to render HTML, it is however a
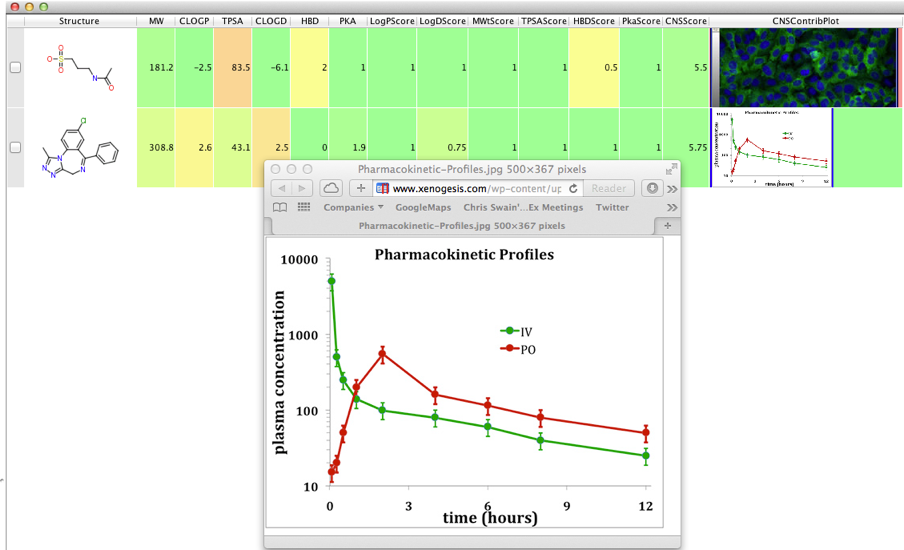
Vortex has a limited capacity to render HTML, it is however a very limited ability so there is no support for javascript or CSS but