Apple have just released OpenELM, a family of Open-source Efficient Language Models. https://huggingface.co/apple/OpenELM
Month: April 2024
The next Cambridge Cheminformatics Meeting – on 8 May 2024, 4pm UK time; Hybrid, at the CCDC and on Zoom! Full details are on the
This looks very useful for anyone having to process multiple molecules, I particularly like the error processing! The open-source package scikit-learn provides various machine learning
tmap is a very fast visualisation library for large, high-dimensional data sets. It was published in 2020 DOI and the code is available on GitHub
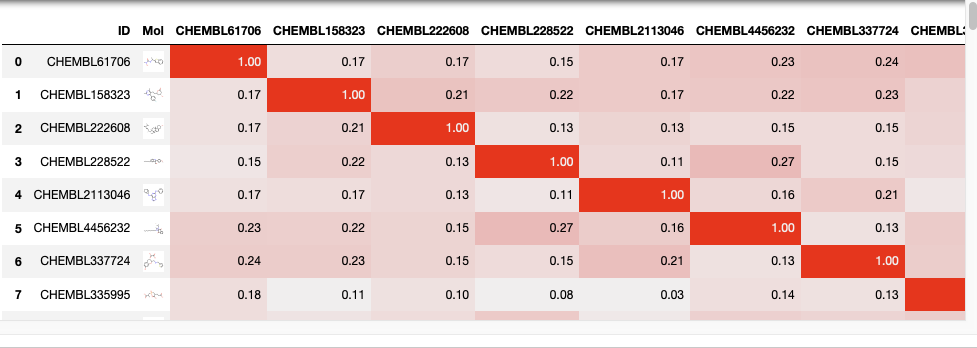
I’m sometimes asked for a tool to compare the similarity of a list of molecules with every other molecule in the list. I suspect there
Interesting paper on Biorxiv https://www.biorxiv.org/content/10.1101/2023.09.27.559736v1.full.pdf We propose a method dedicated to finding antibodies in cryo-EM densities : CrAI. This machine learning method leverages the conserved structure
The Apple worldwide developer conference is on June 10–14, 2024. Whilst there is likely to be updates to Apple hardware and OS, given the recent
A new Apple preprint has appeared on Arxiv. https://arxiv.org/pdf/2403.20329.pdf Reference resolution is an important problem, one that is essential to understand and success- fully handle