Sometimes target identification studies can just turn up a list of Uniprot IDs, whilst there are a number of places you can go to find
Category: Vortex Scripts
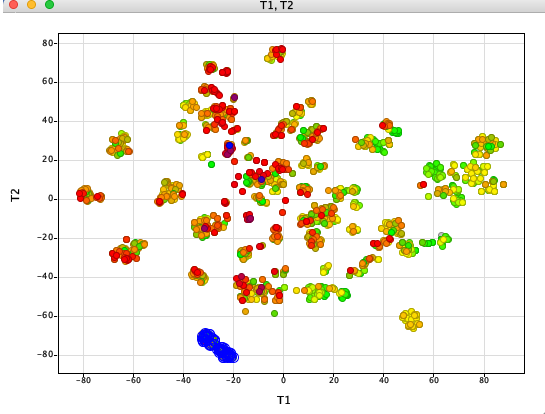
t-distributed stochastic neighbor embedding (t-SNE) is a statistical method for visualizing high-dimensional data by giving each datapoint a location in a two or three-dimensional map. Whilst there are
Prediction of the metabolism of small molecules is very challenging and so having a variety of different tools is always useful. I’ve previously written Vortex
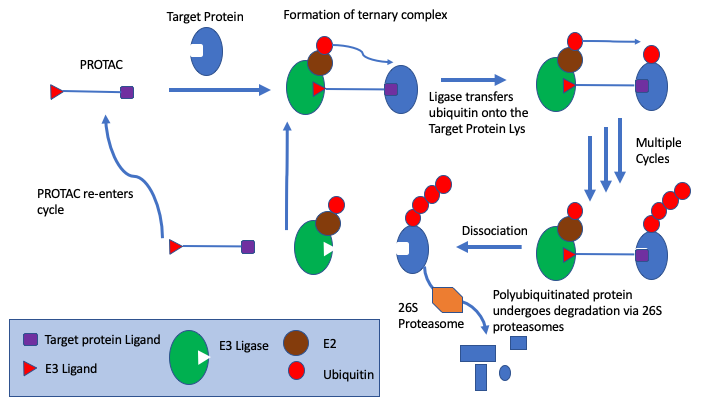
PROteolysis TArgeting Chimeras (PROTACs) technology provides an alternative to module biological function by specially using the ubiquitin proteasome system to induce degradation of the target
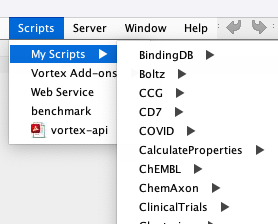
At least on a Mac the default place to store Vortex scripts is in the vortex folder ~/vortex/scripts Any folder/scripts you put in the vortex/scripts
One of the frequent situations after running a screen is you have a list of hits and you want to select related analogues to explore
When making selections from large datasets it is worth mentioning that as datasets get larger a simple random selection is often the best (and quickest)
Sometimes when you import a dataset into a Vortex workspace the default display can be not ideal. For example I imported this CDK7 dataset from
JDBC, which stands for Java Database Connectivity, is a Java API that allows Java applications to interact with relational databases. There are JDBC drivers for most
The previous scripts have allowed the user to get more information about a PDB entry and to import the ligand structures. This script allows the