For a long time the computational chemistry and molecular modelling package MOE from Chemical Computing Group has required XQuartz to provide the windowing environment. With
Category: Hints and Tutorials
Tutorials, Hints, Tips and Tricks
I’ve written about TabPFN previously https://macinchem.org/2025/02/06/looking-at-tabpfn/ and I see a technical report has just been published. https://priorlabs.ai/technical-reports/tabpfn-3 TabPFN is a foundation model trained on around 130,000,000
https://www.linkedin.com/feed/update/urn:li:activity:7461059877155631105 A fantastic step-by-step breakdown of an example Python pipeline used to build a ligand-based 3D pharmacophore model using free, open-source tools.
I’ve written several posts on the various options for clustering molecules https://macinchem.org/?s=clustering and a recent post from NVIDIA described GPU-Accelerated Clustering with nvMolKit that uses
Sometimes target identification studies can just turn up a list of Uniprot IDs, whilst there are a number of places you can go to find
Recently a guest post from NVIDIA described GPU-Accelerated Clustering with nvMolKit that uses CUDA. A recent post no describes a port of the nvMolKit (CUDA) molecular clustering
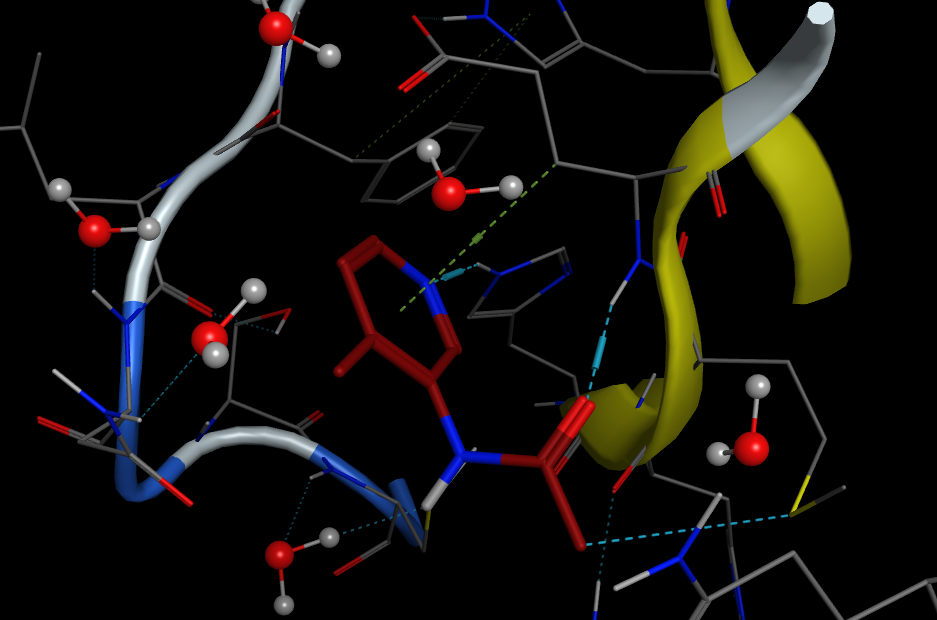
In 2020 as a result of lockdown I was asked to help create a course for MRes students as an introduction to computer-aided drug design.
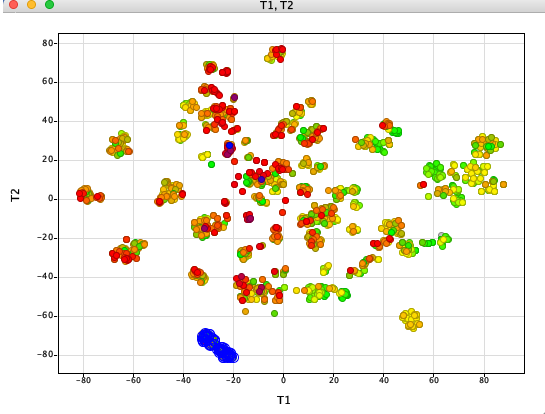
t-distributed stochastic neighbor embedding (t-SNE) is a statistical method for visualizing high-dimensional data by giving each datapoint a location in a two or three-dimensional map. Whilst there are
An interesting web app that fetches ChEMBL bioactivity data for a target (via UniProt ID), computes molecular descriptors, and trains a simple predictive model (regression, with
Prediction of the metabolism of small molecules is very challenging and so having a variety of different tools is always useful. I’ve previously written Vortex