I was recently asked if I could generate IUPAC names for a series of molecules for a patent filing. There are many chemical drawing packages that will generate the IUPAC for a single compound and whilst I could have spent several hours cutting and pasting I decided to write a simple Vortex script to do the task.
The command line tool cxcalc from ChemAxon is the command line version of ChemAxon’s Calculator Plugins, and can be used to calculate a wide variety of properties including the IUPAC name.
The command has the following syntax:
cxcalc [general options] [input file(s)/string(s)] <plugin> [plugin option(s)] [input file(s)/string(s)]
And so to calculate the name the command would be (where sdfFile is the path to the sdf file containing the structures.
/Applications/MarvinSuite/bin/cxcalc', sdfFile, 'name'
As shown below
/Applications/MarvinSuite/bin/cxcalc /Users/username/Desktop/SampleFiles/emend3D.sdf name
id Preferred IUPAC Name
1 5-{[(2R,3S)-2-[(1R)-1-[3,5-bis(trifluoromethyl)phenyl]ethoxy]-3-(4-fluorophenyl)morpholin-4-yl]methyl}-2,3-dihydro-1H-1,2,4-triazol-3-one
We can use subprocess to run the command and capture the output.
The Vortex Script
# Created by http://www.macinchem.org
# Generates IUPAC names using ChemAxon cxcalc
import sys
import subprocess
# Get the path to the currently open sdf file
sdfFile = vortex.getFileForPropertyCalculation(vtable)
# http://www.chemaxon.com/
# Run cxcalc on the file
p = subprocess.Popen(['/Applications/MarvinSuite/bin/cxcalc', sdfFile, 'name'], stdout=subprocess.PIPE)
output = p.communicate()[0]
# Create new columns in table
lines = output.split('\n')
colName = lines[0].split('\t')
for c in colName:
column = vtable.findColumnWithName(c, 1, 3)
vtable.fireTableStructureChanged()
keys = []
for i in lines:
words = i.split('\t')
if len(words) == 2:
keys.append(words[0])
# Parse the output
rows = lines[1:len(lines)]
for r in range(0, vtable.getRealRowCount()):
vals = rows[r].split('\t')
for j in range(0, len(vals)):
column = vtable.findColumnWithName(colName[j], 0)
column.setValueFromString(r, vals[j])
vtable.fireTableStructureChanged()
The result is shown below
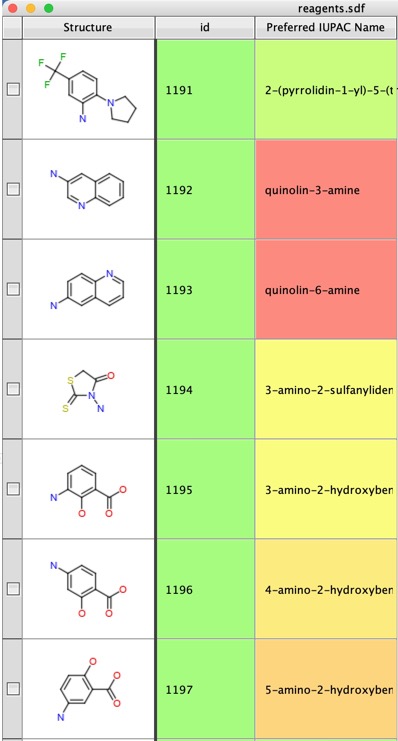
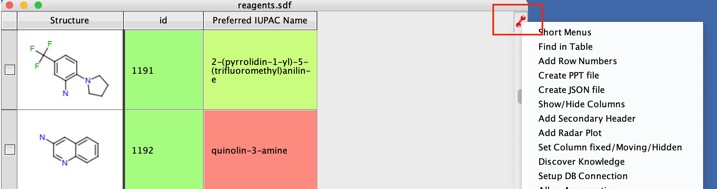
To get the names to wrap with the cell you need to open the “Look and Feel” dialog by clicking on the icon at the top right corner of the table view..
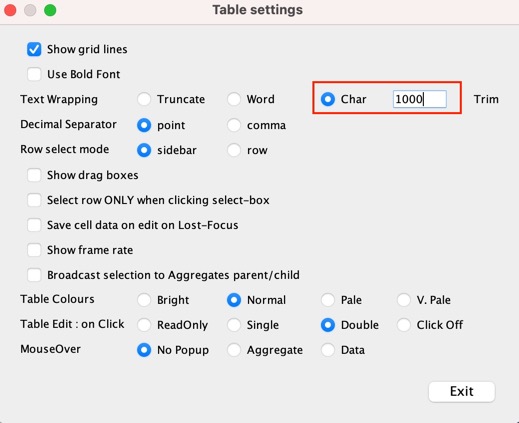
Then edit the Text wrapping options.
Last updated 21 Oct 2021