Lack of data has hampered the building of models to accurately predict binding affinity so I’m sure everyone is super excited to see the first
Tag: compchem
A recent paper on ChemRxiv https://chemrxiv.org/doi/10.26434/chemrxiv-2025-9c1v6 describes ANNalog a transformer-based sequence-to-sequence generative model trained on pairs of molecules extracted from the same bioactivity assay in
Cambridge Cheminformatics Meeting Cambridge Cheminformatics Meeting, 22 April 2026 *** AI Agents for Omics *** Chemical Space Docking for Billion-Sized Libraries *** Quantum Chemistry for
I’ve previously written about mlxmolkit in particular for clustering where see a 40-fold improvement in speed. The toolkit has now been extended to include 3D
Avogadro is a free and open source molecular editor and visualization tool. The new release of Avogadro 2, is a major update, and includes major
I’ve written several posts on the various options for clustering molecules https://macinchem.org/?s=clustering and a recent post from NVIDIA described GPU-Accelerated Clustering with nvMolKit that uses
I’ve just signed up for the meeting, hope to see you there at Burlington House, London, UK Wednesday 8th April 2026 A one-day conference on Chemical
OpenFold3-preview is a biomolecular structure prediction model aiming to be a bitwise reproduction of DeepMind’sAlphaFold3, developed by the AlQuraishi Lab at Columbia University and the
ChemeleonSMD distills the CheMeleon pretrained Directed Message Passing Neural Network (DMPNN) into a SCORE-style architecture running natively on MLX. The SCORE Euler skip connection replaces ChemProp’s hard-overwrite message passing
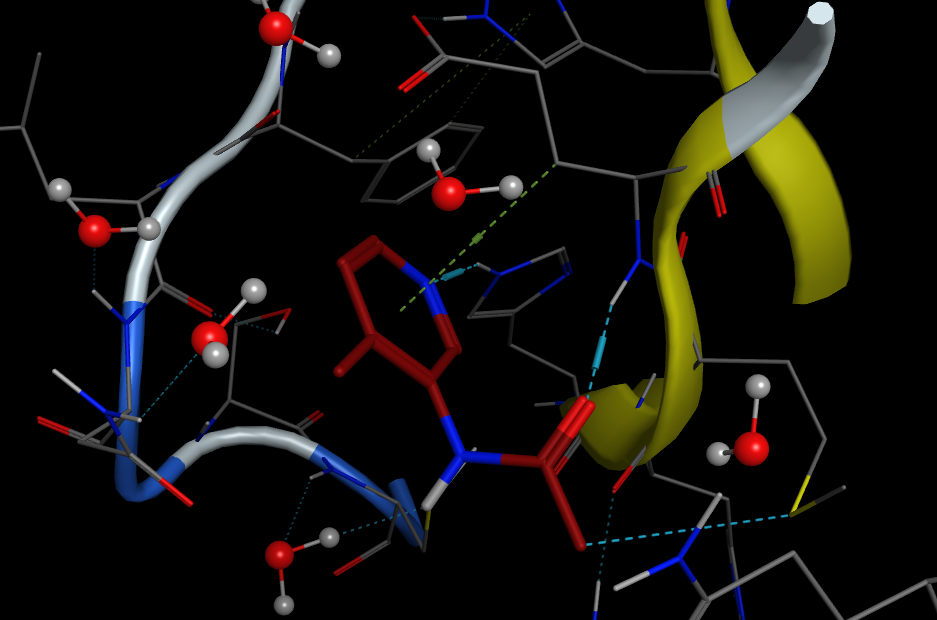
In 2020 as a result of lockdown I was asked to help create a course for MRes students as an introduction to computer-aided drug design.