Sometimes target identification studies can just turn up a list of Uniprot IDs, whilst there are a number of places you can go to find out more information I find ChEMBL is an invaluable source of information. These Vortex scripts use the Uniprot ID to fetch a variety of information.
Uniprot ID
P04058
P22734
P00760
P15207
Q08881
P00742
Q9Y233
O14965
First import the above Uniprot ID into Vortex.
Uniprot to ChEMBL target data Vortex script
To access the ChEMBL data we can use one of the web services, in this case a URL of this format, where the result is returned in json format.
|
1 |
ttps://www.ebi.ac.uk/chembl/api/data/target?target_components__accession=Q13936&format=json |
The Vortex script
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 |
#ChEMBL Targets Target Search Search using uniprotID # Standard imports # Python imports import urllib2 import urllib from com.xhaus.jyson import JysonCodec as json # Vortex imports import com.dotmatics.vortex.util.Util as Util import com.dotmatics.vortex.mol2img.jni.genImage as genImage import com.dotmatics.vortex.mol2img.Mol2Img as mol2Img import jarray import binascii import string import os input_label = swing.JLabel("Uniprot column (for input)") input_cb = workspace.getColumnComboBox() panel = swing.JPanel() layout.fill(panel, input_label, 0, 0) layout.fill(panel, input_cb, 1, 0) ret = vortex.showInDialog(panel, "Choose uniprot column") if ret == vortex.OK: input_idx = input_cb.getSelectedIndex() if input_idx == 0: vortex.alert("you must choose a column") else: col = vtable.getColumn(input_idx - 1) rows = vtable.getRealRowCount() for r in range(0, int(rows)): uniprotID = col.getValueAsString(r) mystr = "https://www.ebi.ac.uk/chembl/api/data/target?target_components__accession=" + uniprotID + "&format=json" try: myreturn = urllib2.urlopen(mystr).read() except urllib2.HTTPError: continue # some not found Target not found for accession:P55957 # if myreturn.find('Target not found') != -1: j = json.loads(myreturn) try: TheData = str(j['targets'][0]['target_chembl_id']) colChemblID = vtable.findColumnWithName('target_chembl_id', 1) colChemblID.setValueFromString(r, TheData) TheData = str(j['targets'][0]['pref_name']) colType = vtable.findColumnWithName('pref_name', 1) colType.setValueFromString(r, TheData) TheData = str(j['targets'][0]['organism']) colType = vtable.findColumnWithName('organism', 1) colType.setValueFromString(r, TheData) TheData = str(j['targets'][0]['target_type']) colType = vtable.findColumnWithName('target_type', 1) colType.setValueFromString(r, TheData) except: colChemblID = vtable.findColumnWithName('target_chembl_id', 1) colChemblID.setValueFromString(r, "Not known") vtable.fireTableStructureChanged() #mystr = "https://www.ebi.ac.uk/chembl/api/data/target?target_components__accession=Q13936&format=json" # data format{"page_meta": {"limit": 20, "next": null, "offset": 0, "previous": null, "total_count": 5}, "targets": [{"cross_references": [{"xref_id": "Cav1.2", "xref_name": null, "xref_src": "Wikipedia"}], "organism": "Homo sapiens", "pref_name": "Voltage-gated L-type calcium channel alpha-1C subunit", "species_group_flag": false, "target_chembl_id": "CHEMBL1940", "target_components": [{"accession": "Q13936", "component_description": "Voltage-dependent L-type calcium channel subunit alpha-1C", "component_id": 261, "component_type": "PROTEIN", "relationship": "SINGLE PROTEIN", "target_component_synonyms": [{"component_synonym": "CACH2", "syn_type": "GENE_SYMBOL_OTHER"}, {"component_synonym": "CACN2", "syn_type": "GENE_SYMBOL_OTHER"}, {"component_synonym": "CACNA1C", "syn_type": "GENE_SYMBOL"}, {"component_synonym": "CACNL1A1", "syn_type": "GENE_SYMBOL_OTHER"}, {"component_synonym": "Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle", "syn_type": "UNIPROT"}, {"component_synonym": "CCHL1A1", "syn_type": "GENE_SYMBOL_OTHER"}, {"component_synonym": "Voltage-dependent L-type calcium channel subunit alpha-1C", "syn_type": "UNIPROT"}, {"component_synonym": "Voltage-gated calcium channel subunit alpha Cav1.2", "syn_type": "UNIPROT"}], "target_component_xrefs": [{"xref_id": "CACNA1C", "xref_name": "Brugada syndrome 3; Timothy syndrome", "xref_src_db": "CGD"}, {"xref_id": "ENSG00000151067", "xref_name": null, "xref_src_db": "EnsemblGene"}, {"xref_id": "ENSG00000285479", "xref_name": null, "xref_src_db": "EnsemblGene"}, {"xref_id": "Q13936", "xref_name": null, "xref_ |
The first part of the script constructs a dialog box for the usr to identify the column containing the Uniprot ID. Then we construct the URL (mystr), and use it to query ChEMBL, the returned json is then parsed to find the appropriate data and populate the columns created, as shown below.
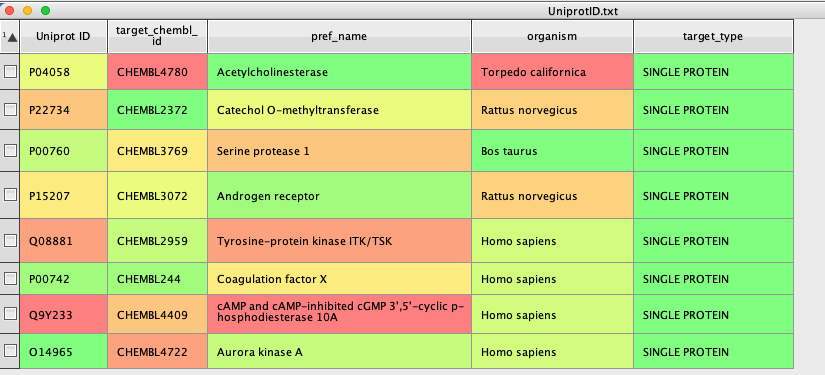
The new columns contain the ChEMBLID for the target, a preferred name, the organism and the target type. The script could be modified to extract additional information.
Vortex script to find bioactivities.
The ChEMBLID now provides an identifier that allows us to query ChEMBL for a variety of information. The URL below will return detailed information about actives stored in ChEMBL.
|
1 |
https://www.ebi.ac.uk/chembl/api/data/activity.json?target_chembl_id=CHEMBL2971 |
The Vortex script
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 |
#How many biioactivities for a CHEMBL target. # Python imports import urllib2 import urllib from com.xhaus.jyson import JysonCodec as json # Vortex imports import com.dotmatics.vortex.util.Util as Util import com.dotmatics.vortex.mol2img.jni.genImage as genImage import com.dotmatics.vortex.mol2img.Mol2Img as mol2Img import jarray import binascii import string import os input_label = swing.JLabel("ChEMBLID column (for input)") input_cb = workspace.getColumnComboBox() panel = swing.JPanel() layout.fill(panel, input_label, 0, 0) layout.fill(panel, input_cb, 1, 0) ret = vortex.showInDialog(panel, "Choose CHEMBLID column") if ret == vortex.OK: input_idx = input_cb.getSelectedIndex() if input_idx == 0: vortex.alert("you must choose a column") else: col = vtable.getColumn(input_idx - 1) rows = vtable.getRealRowCount() for r in range(0, int(rows)): ChEMBLID = col.getValueAsString(r) mystr = "https://www.ebi.ac.uk/chembl/api/data/activity.json?target_chembl_id=" + ChEMBLID try: myreturn = urllib2.urlopen(mystr).read() except urllib2.HTTPError: continue # some not found Target not found for accession:P55957 # if myreturn.find('Target not found') != -1: j = json.loads(myreturn) try: TheData =str(j['page_meta']['total_count']) colNumActives = vtable.findColumnWithName('Num Bioactivities', 1) colNumActives.setValueFromString(r, TheData) except: colNumActives.setValueFromString(r, "Not known") vtable.fireTableStructureChanged() #mystr = "https://www.ebi.ac.uk/chembl/api/data/activity.json?target_chembl_id=CHEMBL2971" |
In a similar manner to the previous script, the first part constructs the dialog box to allow the user to identify the column containing the ChEMBLID. Then we construct the URL (mystr), and use it to query ChEMBL, the returned json is then parsed to find the appropriate data “total_count” and populate the column created, as shown below.
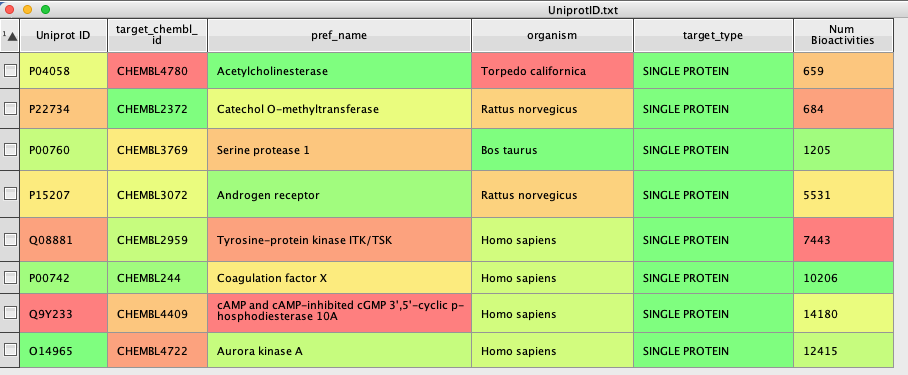
The returned json actually contains much more information. It is possible to download all data from a specific ChEMBL web service resource. This is made possible by returning responses from the web services in ‘pages’, which can be navigated through using a ‘page_meta’ section. The ‘page_meta’ section includes information about total number of hits, total number of pages and links to the next and previous pages. An example ‘page_meta’ section is displayed below.
|
1 2 3 4 5 6 7 8 9 |
"page_meta": { "limit": 20, "next": "/chembl/api/data/activity.json?limit=20&offset=20", "offset": 0, "previous": null, "total_count": 13520737 } |
For this script we only use the last element of the json. If you want to access all the information you will need to use the page_meta section to navigate to subsequent pages.
The scripts can be downloaded here
